Nice find! Here's a bit more information:
ERVs, a startling line of evidence
from genetics (not fossils) provides additional, overwhelming support for human/chimp common ancestry, one that establishes it beyond a shadow of a doubt.
Much like stellar parallax could be considered 'proof' of heliocentricism, so too do unique fingerprints of ancient retroviral infection demonstrate relatedness if found in the genome of two different organisms.
1. Background
Viral infections. We all know and hate them. From the common cold to the common cold
sore, they're annoying little buggers. The virus particle invades a cell, injects its proprietary genome, hijacks the cellular machinery and turns it into a virus-making factory. Upon successful completion of this process, the cell begins releasing fresh copies of the virus, often by bursting and dying.
Occasionally, however, something goes wrong in the integration process. When this occurs,
the cell can survive, but
selected portions of the viral DNA remain in its genome. These vary from
infection to infection, not just from virus to virus.
In effect, the cell becomes permanently scarred and
carries the unique, identifiable fingerprint of its assailant. (
Source)
Since the markers are now in the genome, they become hereditary--hence, any descendant of the afflicted cell will contain the same exact viral fingerprint(s) at the same exact spot in its DNA.
Viral infections are, of course, not limited to the mucous membranes, blood stream and lungs. Under certain conditions, they can also infect a germ line (egg or sperm) cell. The odds of this cell now surviving to adulthood are astronomically low; not only does it have to survive a viral infection, but be one of the extremely lucky few that ever make it to merge with their counterpart and form a viable zygote.
However, this does occur. When it does, the resulting offspring has the unique viral fingerprint embedded in every cell in its body, what is known as an "endogenous retroviral insertion", henceforth referred to as an ERV. (
Source)
ERVs can be easily identified by their extreme similarity to known retroviruses. They even contain vestiges of
gag,
env and
pol genes that code for viral surface proteins (
explanation of GEP protein coats,
information on their presence in ERVs).
We've directly observed creation of new, identifiable ERVs by viral integration into cells
in vitro, as well, so this process is as well-documented (if not more) as fossil formation. When it occurs in ordinary cells, it can cripple vital genes, subsequently causing cancer. High-risk papillomaviruses have been indicted as one of the leading causes of cervical tumors, for example. (
Source)
Like any genes, the viral elements can then piggyback on the success of an individual and become established in the entire species. In small populations, this can easily occur via
genetic drift (random establishment of selectively neutral or deleterious alleles). In larger ones, natural selection can work to establish the few ERVs that yield a benefit to their host.
Each step in the process is not very common, and has unique aspects that are extremely unlikely to be reproduced in two independent cases of ERV creation. Hence, when they all occur, the organisms possessing the ERV are marked with a molecular fossil that will prove indispensible in tracing their origin.
2. Evolutionary Implications of ERVs
In a nutshell, this can serve as a "paternity test" of sorts: if you find the same endogenous retroviral insertion in two different organisms, the only viable explanation is that they inherited same from a common ancestor that, itself, possessed it. Offshoot species will have the same unlikely and easily identifiable ERVs, enabling us to construct accurate phylogenies from an independent line of evidence.
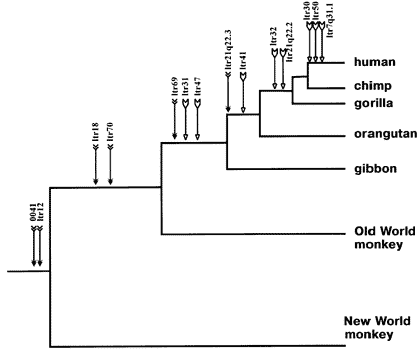
Human endogenous retroviral insertions in identical chromosomal locations in various primates. Notice just how well the standard evolutionary phylogeny, derived from fossil and anatomical evidence (humans and chimps closest, then orangutans and gorillas, then gibbons, then old world monkeys, then new world monkeys) is corroborated by this uniquely independent line of evidence.(
Source)
As a side note, ERVs have also been used to reconstruct the relationships between dogs, jackals, wolves and foxes, among other species of domestic animal.
Needless to say, this offers a quick and easy way to experimentally test if any two similar organisms are
unrelated. If they are, they won't share any ERVs. If you find the same one at the same place in their genomes, however, the hypothesis they originated independently is effectively falsified.
Due to the nature of the branching tree of life, ERVs shared between distantly-related organisms
will have made their initial appearance in their common ancestor. Hence, they would have also been present as
further, closer organisms split off from the lineage; if you find an ERV common to humans and monkeys, the ancestor of chimps must have had the insertion as well. Similarly, ERVs found in New World Monkeys and chimps will usually be present in humans. ERVs found in humans and gorillas should be found in chimps. And so on. This provides an additional way to test evolutionary predictions.
It should be noted that there are rare but observable genetic phenomena that can remove ERVs from the genome. However, we can almost always
tell when this has occured, such as identifying the characteristic fingerprint of a deletion mutation right before the ERV 'should have' been found.
Nevertheless, the more basic question of '
are there shared ERVs between these species
at all?' is a much more stringent, rock-solid test of evolution than the specific phylogenetic relationships
individual insertions indicate, which can be erroneous on occasion.
When analyzed as a statistical whole, ERV insertion patterns will also recapitulate phylogenies drawn from independent data like the fossil record and comparative anatomy. This is a remarkable confirmation, and totally at odds with the predictions made by the hypothesis of independent origin of species.
As you can see by now, this is the strongest support for evolution I've ever come across; a truly powerful and damning smoking gun.