-
Starting today August 7th, 2024, in order to post in the Married Couples, Courting Couples, or Singles forums, you will not be allowed to post if you have your Marital status designated as private. Announcements will be made in the respective forums as well but please note that if yours is currently listed as Private, you will need to submit a ticket in the Support Area to have yours changed.
-
CF has always been a site that welcomes people from different backgrounds and beliefs to participate in discussion and even debate. That is the nature of its ministry. In view of recent events emotions are running very high. We need to remind people of some basic principles in debating on this site. We need to be civil when we express differences in opinion. No personal attacks. Avoid you, your statements. Don't characterize an entire political party with comparisons to Fascism or Communism or other extreme movements that committed atrocities. CF is not the place for broad brush or blanket statements about groups and political parties. Put the broad brushes and blankets away when you come to CF, better yet, put them in the incinerator. Debate had no place for them. We need to remember that people that commit acts of violence represent themselves or a small extreme faction.
You are using an out of date browser. It may not display this or other websites correctly.
You should upgrade or use an alternative browser.
You should upgrade or use an alternative browser.
The phenomenon and the explanation
- Thread starter Phred
- Start date
- Jan 9, 2018
- 3,132
- 871
- Country
- United States
- Gender
- Male
- Faith
- Christian
- Marital Status
- Single
More precisely, intellect is not located.Umm. The brain.
There is no intellect located within any other part of the human.
Upvote
0
Hans Blaster
I march with Sherman
- Mar 11, 2017
- 23,201
- 17,247
- 55
- Country
- United States
- Gender
- Male
- Faith
- Atheist
- Marital Status
- Private
- Politics
- US-Democrat
More precisely, intellect is not located.
No that's less precise, it is just babble. What on Earth do you mean by this?
Upvote
0
- Jan 9, 2018
- 3,132
- 871
- Country
- United States
- Gender
- Male
- Faith
- Christian
- Marital Status
- Single
Has anyone located the organ from which the knowledge and freedom to do right or wrong comes from? How about the guilt and reward felt doing it? If not then the source of the power to do so has no location.No that's less precise, it is just babble. What on Earth do you mean by this?
Upvote
0
Hans Blaster
I march with Sherman
- Mar 11, 2017
- 23,201
- 17,247
- 55
- Country
- United States
- Gender
- Male
- Faith
- Atheist
- Marital Status
- Private
- Politics
- US-Democrat
Has anyone located the organ from which the knowledge and freedom to do right or wrong comes from? How about the guilt and reward felt doing it? If not then the source of the power to do so has no location.
Those are all thoughts. Thoughts arise from the brain. Have you left yours on the night stand?
Upvote
0
- Jan 9, 2018
- 3,132
- 871
- Country
- United States
- Gender
- Male
- Faith
- Christian
- Marital Status
- Single
Just thoughts? Huh. Free Will?Those are all thoughts. Thoughts arise from the brain. Have you left yours on the night stand?
Ok. Personal experience from childhood.
When I was eight I wanted to hunt. I got a bb gun for my birthday. I didn't want to shoot a target I wanted to hunt. I decided to hunt for bird. A bird sitting on the lowest wire coming from the telephone pole. I aimed ... fired, plop, the bird tumbled to the ground and landed as if sitting in a chair. It's head stretched up looking to and fro screeching. Right then I was pounded with shame. It hunched me over and I scurried to a bush hunched over to hide. It came in waves. took me time to go inside the house to recover.
Where did that shame come from?
Last edited:
Upvote
0
Your brain.Just thoughts? Huh. Free Will?
Ok. Personal experience from childhood.
When I was eight I wanted to hunt. I got a bb gun for my birthday. I didn't want to shoot a target I wanted to hunt. I decided to hunt for bird. A bird sitting on the lowest wire coming from the telephone pole. I aimed ... fired, plop, the bird tumbled to the ground and landed as if sitting in a chair. It's head stretched up looking to and fro screeching. Right then I was pounded with shame. It hunched me over and I scurried to a bush hunched over to hide. It came in waves. took me time to go inside the house to recover.
Where did that shame come from?
Upvote
0
tas8831
Well-Known Member
- May 5, 2017
- 5,611
- 3,999
- 56
- Country
- United States
- Gender
- Male
- Faith
- Atheist
- Marital Status
- Married
A shame that @Mark Quayle could not/would not address any of this in any substantive way. Even more of a shame that he keeps ignoring that which he claims - like so many creationists - is all about bias and such. Typical, really.Evolution is your claim.
=======
Clearly - you ignored it both [added in edit: all three] times I posted it for you - the first time, you merely omitted it from your reply. No, you just ignored it - which I guess you feel you must do in order to keep believing your inflammatory and unsupported charges of circular reasoning and the like. Third time a charm?
-------
I forget now who originally posted these on this forum, but I keep it in my archives because it offers a nice 'linear' progression of testing a methodology and then applying it.
The tested methodology:
Science 25 October 1991:
Vol. 254. no. 5031, pp. 554 - 558
Gene trees and the origins of inbred strains of mice
WR Atchley and WM Fitch
Extensive data on genetic divergence among 24 inbred strains of mice provide an opportunity to examine the concordance of gene trees and species trees, especially whether structured subsamples of loci give congruent estimates of phylogenetic relationships. Phylogenetic analyses of 144 separate loci reproduce almost exactly the known genealogical relationships among these 24 strains. Partitioning these loci into structured subsets representing loci coding for proteins, the immune system and endogenous viruses give incongruent phylogenetic results. The gene tree based on protein loci provides an accurate picture of the genealogical relationships among strains; however, gene trees based upon immune and viral data show significant deviations from known genealogical affinities.
======================
Science, Vol 255, Issue 5044, 589-592
Experimental phylogenetics: generation of a known phylogeny
DM Hillis, JJ Bull, ME White, MR Badgett, and IJ Molineux
Department of Zoology, University of Texas, Austin 78712.
Although methods of phylogenetic estimation are used routinely in comparative biology, direct tests of these methods are hampered by the lack of known phylogenies. Here a system based on serial propagation of bacteriophage T7 in the presence of a mutagen was used to create the first completely known phylogeny. Restriction-site maps of the terminal lineages were used to infer the evolutionary history of the experimental lines for comparison to the known history and actual ancestors. The five methods used to reconstruct branching pattern all predicted the correct topology but varied in their predictions of branch lengths; one method also predicts ancestral restriction maps and was found to be greater than 98 percent accurate.
==================================
Science, Vol 264, Issue 5159, 671-677
Application and accuracy of molecular phylogenies
DM Hillis, JP Huelsenbeck, and CW Cunningham
Department of Zoology, University of Texas, Austin 78712.
Molecular investigations of evolutionary history are being used to study subjects as diverse as the epidemiology of acquired immune deficiency syndrome and the origin of life. These studies depend on accurate estimates of phylogeny. The performance of methods of phylogenetic analysis can be assessed by numerical simulation studies and by the experimental evolution of organisms in controlled laboratory situations. Both kinds of assessment indicate that existing methods are effective at estimating phylogenies over a wide range of evolutionary conditions, especially if information about substitution bias is used to provide differential weightings for character transformations.
Application of the tested methodology:
Implications of natural selection in shaping 99.4% nonsynonymous DNA identity between humans and chimpanzees: Enlarging genus Homo
"Here we compare ≈90 kb of coding DNA nucleotide sequence from 97 human genes to their sequenced chimpanzee counterparts and to available sequenced gorilla, orangutan, and Old World monkey counterparts, and, on a more limited basis, to mouse. The nonsynonymous changes (functionally important), like synonymous changes (functionally much less important), show chimpanzees and humans to be most closely related, sharing 99.4% identity at nonsynonymous sites and 98.4% at synonymous sites. "
Mitochondrial Insertions into Primate Nuclear Genomes Suggest the Use of numts as a Tool for Phylogeny
"Moreover, numts identified in gorilla Supercontigs were used to test the human–chimp–gorilla trichotomy, yielding a high level of support for the sister relationship of human and chimpanzee."
A Molecular Phylogeny of Living Primates
"Once contentiously debated, the closest human relative of chimpanzee (Pan) within subfamily Homininae (Gorilla, Pan, Homo) is now generally undisputed. The branch forming the Homo andPanlineage apart from Gorilla is relatively short (node 73, 27 steps MP, 0 indels) compared with that of thePan genus (node 72, 91 steps MP, 2 indels) and suggests rapid speciation into the 3 genera occurred early in Homininae evolution. Based on 54 gene regions, Homo-Pan genetic distance range from 6.92 to 7.90×10−3 substitutions/site (P. paniscus and P. troglodytes, respectively), which is less than previous estimates based on large scale sequencing of specific regions such as chromosome 7[50]. "
Catarrhine phylogeny: noncoding DNA evidence for a diphyletic origin of the mangabeys and for a human-chimpanzee clade.
"The Superfamily Hominoidea for apes and humans is reduced to family Hominidae within Superfamily Cercopithecoidea, with all living hominids placed in subfamily Homininae; and (4) chimpanzees and humans are members of a single genus, Homo, with common and bonobo chimpanzees placed in subgenus H. (Pan) and humans placed in subgenus H. (Homo). It may be noted that humans and chimpanzees are more than 98.3% identical in their typical nuclear noncoding DNA and probably more than 99.5% identical in the active coding nucleotide sequences of their functional nuclear genes (Goodman et al., 1989, 1990). In mammals such high genetic correspondence is commonly found between sibling species below the generic level but not between species in different genera."
The tested methodology:
Science 25 October 1991:
Vol. 254. no. 5031, pp. 554 - 558
Gene trees and the origins of inbred strains of mice
WR Atchley and WM Fitch
Extensive data on genetic divergence among 24 inbred strains of mice provide an opportunity to examine the concordance of gene trees and species trees, especially whether structured subsamples of loci give congruent estimates of phylogenetic relationships. Phylogenetic analyses of 144 separate loci reproduce almost exactly the known genealogical relationships among these 24 strains. Partitioning these loci into structured subsets representing loci coding for proteins, the immune system and endogenous viruses give incongruent phylogenetic results. The gene tree based on protein loci provides an accurate picture of the genealogical relationships among strains; however, gene trees based upon immune and viral data show significant deviations from known genealogical affinities.
======================
Science, Vol 255, Issue 5044, 589-592
Experimental phylogenetics: generation of a known phylogeny
DM Hillis, JJ Bull, ME White, MR Badgett, and IJ Molineux
Department of Zoology, University of Texas, Austin 78712.
Although methods of phylogenetic estimation are used routinely in comparative biology, direct tests of these methods are hampered by the lack of known phylogenies. Here a system based on serial propagation of bacteriophage T7 in the presence of a mutagen was used to create the first completely known phylogeny. Restriction-site maps of the terminal lineages were used to infer the evolutionary history of the experimental lines for comparison to the known history and actual ancestors. The five methods used to reconstruct branching pattern all predicted the correct topology but varied in their predictions of branch lengths; one method also predicts ancestral restriction maps and was found to be greater than 98 percent accurate.
==================================
Science, Vol 264, Issue 5159, 671-677
Application and accuracy of molecular phylogenies
DM Hillis, JP Huelsenbeck, and CW Cunningham
Department of Zoology, University of Texas, Austin 78712.
Molecular investigations of evolutionary history are being used to study subjects as diverse as the epidemiology of acquired immune deficiency syndrome and the origin of life. These studies depend on accurate estimates of phylogeny. The performance of methods of phylogenetic analysis can be assessed by numerical simulation studies and by the experimental evolution of organisms in controlled laboratory situations. Both kinds of assessment indicate that existing methods are effective at estimating phylogenies over a wide range of evolutionary conditions, especially if information about substitution bias is used to provide differential weightings for character transformations.
Application of the tested methodology:
Implications of natural selection in shaping 99.4% nonsynonymous DNA identity between humans and chimpanzees: Enlarging genus Homo
"Here we compare ≈90 kb of coding DNA nucleotide sequence from 97 human genes to their sequenced chimpanzee counterparts and to available sequenced gorilla, orangutan, and Old World monkey counterparts, and, on a more limited basis, to mouse. The nonsynonymous changes (functionally important), like synonymous changes (functionally much less important), show chimpanzees and humans to be most closely related, sharing 99.4% identity at nonsynonymous sites and 98.4% at synonymous sites. "
Mitochondrial Insertions into Primate Nuclear Genomes Suggest the Use of numts as a Tool for Phylogeny
"Moreover, numts identified in gorilla Supercontigs were used to test the human–chimp–gorilla trichotomy, yielding a high level of support for the sister relationship of human and chimpanzee."
A Molecular Phylogeny of Living Primates
"Once contentiously debated, the closest human relative of chimpanzee (Pan) within subfamily Homininae (Gorilla, Pan, Homo) is now generally undisputed. The branch forming the Homo andPanlineage apart from Gorilla is relatively short (node 73, 27 steps MP, 0 indels) compared with that of thePan genus (node 72, 91 steps MP, 2 indels) and suggests rapid speciation into the 3 genera occurred early in Homininae evolution. Based on 54 gene regions, Homo-Pan genetic distance range from 6.92 to 7.90×10−3 substitutions/site (P. paniscus and P. troglodytes, respectively), which is less than previous estimates based on large scale sequencing of specific regions such as chromosome 7[50]. "
Catarrhine phylogeny: noncoding DNA evidence for a diphyletic origin of the mangabeys and for a human-chimpanzee clade.
"The Superfamily Hominoidea for apes and humans is reduced to family Hominidae within Superfamily Cercopithecoidea, with all living hominids placed in subfamily Homininae; and (4) chimpanzees and humans are members of a single genus, Homo, with common and bonobo chimpanzees placed in subgenus H. (Pan) and humans placed in subgenus H. (Homo). It may be noted that humans and chimpanzees are more than 98.3% identical in their typical nuclear noncoding DNA and probably more than 99.5% identical in the active coding nucleotide sequences of their functional nuclear genes (Goodman et al., 1989, 1990). In mammals such high genetic correspondence is commonly found between sibling species below the generic level but not between species in different genera."
CONCLUSION:
This evidence lays out the results of employing a tested methodology on the question of Primate evolution. The same general criteria/methods have been used on nearly all facets of the evolution of living things.
As you like to pretend you understand all this, I eagerly await your analysis in which you point out the confirmation bias, circular reasoning, etc. that you pretend all actual science is based on.
But I will not be holding my breath. All hat, no cattle?
+++++++++
And he never did....
Upvote
0
- May 28, 2018
- 14,362
- 6,414
- 69
- Country
- United States
- Gender
- Male
- Faith
- Reformed
- Marital Status
- Widowed
As you like to pretend you understand all this, I eagerly await your analysis in which you point out the confirmation bias, circular reasoning, etc. that you pretend all actual science is based on.
Where did I claim I understand all this? Since apparently I have not been clear, I don't understand all the terminology, but when I read, either stated or implied, "could be", "suggests", "probably", "this would also work if", and so on, I can see what is happening there.
There is also, "we know that...", when philosophy of science itself demands that there be no final conclusions. Maybe we can regard some things as trustworthy for now, but concerning which a decent bit of skepticism is prudent, but that isn't what (it seems to me) the whole house of cards is built on.
Meanwhile, you have failed to convince me of your positive claim of your brand of evolution.
Upvote
0
tas8831
Well-Known Member
- May 5, 2017
- 5,611
- 3,999
- 56
- Country
- United States
- Gender
- Male
- Faith
- Atheist
- Marital Status
- Married
You claimed that it was all circular reasoning and all that garbage.Where did I claim I understand all this?
Did you forget your vacuous, pompous, unwarranted and wholly unsupported assertions?
Yet you think you can (and have) somehow reject all of the science despite not being able to understand it. Typical hubris of the creationist.Since apparently I have not been clear, I don't understand all the terminology,
Implied? wow... Your posts clearly IMPLY that you have a teenager's understanding of the relevant science, and use this naivete as a cudgel to prop up your fantasies.but when I read, either stated or implied, "could be", "suggests", "probably", "this would also work if", and so on, I can see what is happening there.
Great. So you are unfamiliar with scientific writing. That means it is all conspiracies and lies how, exactly?
So you can't understand the science, but still feel you can dismiss it all. TypicalThere is also, "we know that...", when philosophy of science itself demands that there be no final conclusions. Maybe we can regard some things as trustworthy for now, but concerning which a decent bit of skepticism is prudent, but that isn't what (it seems to me) the whole house of cards is built on.
Meanwhile, you have admitted that you did not even read it, much less understand it.Meanwhile, you have failed to convince me of your positive claim of your brand of evolution.
Typical dishonest creationist, desperate to prop up a failing faith. Here is how I know you are dishonest and that your word-parsing creationist antics keep you ignorant. Regarding the links/quotes below:
1. The phrase "could be" does not appear once.
2. The word "probably" appears once:
"It may be noted that humans and chimpanzees are more than 98.3% identical in their typical nuclear noncoding DNA and probably more than 99.5% identical in the active coding nucleotide sequences of their functional nuclear genes (Goodman et al., 1989, 1990)."
How does that prop up your desperate, uninformed creationism again? How does that negate or diminish the findings when the 'probably' refers to the work of others?
3. The word "suggests" appears once:
"The branch forming the Homo and Pan lineage apart from Gorilla is relatively short (node 73, 27 steps MP, 0 indels) compared with that of the Pan genus (node 72, 91 steps MP, 2 indels) and suggests rapid speciation into the 3 genera occurred early in Homininae evolution. "
How does that help your cause again?
4. The dopey phrase "this would also work if" does not appear anywhere, of course.
You are out of your depth, and in your desperation, continue to make yourself look disingenuous and dishonest. Keep up the good work - another one to refer people to check out as an example of how creationists operate.
I forget now who originally posted these on this forum, but I keep it in my archives because it offers a nice 'linear' progression of testing a methodology and then applying it.
The tested methodology:
Science 25 October 1991:
Vol. 254. no. 5031, pp. 554 - 558
Gene trees and the origins of inbred strains of mice
WR Atchley and WM Fitch
Extensive data on genetic divergence among 24 inbred strains of mice provide an opportunity to examine the concordance of gene trees and species trees, especially whether structured subsamples of loci give congruent estimates of phylogenetic relationships. Phylogenetic analyses of 144 separate loci reproduce almost exactly the known genealogical relationships among these 24 strains. Partitioning these loci into structured subsets representing loci coding for proteins, the immune system and endogenous viruses give incongruent phylogenetic results. The gene tree based on protein loci provides an accurate picture of the genealogical relationships among strains; however, gene trees based upon immune and viral data show significant deviations from known genealogical affinities.
======================
Science, Vol 255, Issue 5044, 589-592
Experimental phylogenetics: generation of a known phylogeny
DM Hillis, JJ Bull, ME White, MR Badgett, and IJ Molineux
Department of Zoology, University of Texas, Austin 78712.
Although methods of phylogenetic estimation are used routinely in comparative biology, direct tests of these methods are hampered by the lack of known phylogenies. Here a system based on serial propagation of bacteriophage T7 in the presence of a mutagen was used to create the first completely known phylogeny. Restriction-site maps of the terminal lineages were used to infer the evolutionary history of the experimental lines for comparison to the known history and actual ancestors. The five methods used to reconstruct branching pattern all predicted the correct topology but varied in their predictions of branch lengths; one method also predicts ancestral restriction maps and was found to be greater than 98 percent accurate.
==================================
Science, Vol 264, Issue 5159, 671-677
Application and accuracy of molecular phylogenies
DM Hillis, JP Huelsenbeck, and CW Cunningham
Department of Zoology, University of Texas, Austin 78712.
Molecular investigations of evolutionary history are being used to study subjects as diverse as the epidemiology of acquired immune deficiency syndrome and the origin of life. These studies depend on accurate estimates of phylogeny. The performance of methods of phylogenetic analysis can be assessed by numerical simulation studies and by the experimental evolution of organisms in controlled laboratory situations. Both kinds of assessment indicate that existing methods are effective at estimating phylogenies over a wide range of evolutionary conditions, especially if information about substitution bias is used to provide differential weightings for character transformations.
Application of the tested methodology:
Implications of natural selection in shaping 99.4% nonsynonymous DNA identity between humans and chimpanzees: Enlarging genus Homo
"Here we compare ≈90 kb of coding DNA nucleotide sequence from 97 human genes to their sequenced chimpanzee counterparts and to available sequenced gorilla, orangutan, and Old World monkey counterparts, and, on a more limited basis, to mouse. The nonsynonymous changes (functionally important), like synonymous changes (functionally much less important), show chimpanzees and humans to be most closely related, sharing 99.4% identity at nonsynonymous sites and 98.4% at synonymous sites. "
Mitochondrial Insertions into Primate Nuclear Genomes Suggest the Use of numts as a Tool for Phylogeny
"Moreover, numts identified in gorilla Supercontigs were used to test the human–chimp–gorilla trichotomy, yielding a high level of support for the sister relationship of human and chimpanzee."
A Molecular Phylogeny of Living Primates
"Once contentiously debated, the closest human relative of chimpanzee (Pan) within subfamily Homininae (Gorilla, Pan, Homo) is now generally undisputed. The branch forming the Homo and Pan lineage apart from Gorilla is relatively short (node 73, 27 steps MP, 0 indels) compared with that of the Pan genus (node 72, 91 steps MP, 2 indels) and suggests rapid speciation into the 3 genera occurred early in Homininae evolution. Based on 54 gene regions, Homo-Pan genetic distance range from 6.92 to 7.90×10−3 substitutions/site (P. paniscus and P. troglodytes, respectively), which is less than previous estimates based on large scale sequencing of specific regions such as chromosome 7[50]. "
Catarrhine phylogeny: noncoding DNA evidence for a diphyletic origin of the mangabeys and for a human-chimpanzee clade.
"The Superfamily Hominoidea for apes and humans is reduced to family Hominidae within Superfamily Cercopithecoidea, with all living hominids placed in subfamily Homininae; and (4) chimpanzees and humans are members of a single genus, Homo, with common and bonobo chimpanzees placed in subgenus H. (Pan) and humans placed in subgenus H. (Homo). It may be noted that humans and chimpanzees are more than 98.3% identical in their typical nuclear noncoding DNA and probably more than 99.5% identical in the active coding nucleotide sequences of their functional nuclear genes (Goodman et al., 1989, 1990). In mammals such high genetic correspondence is commonly found between sibling species below the generic level but not between species in different genera."
The tested methodology:
Science 25 October 1991:
Vol. 254. no. 5031, pp. 554 - 558
Gene trees and the origins of inbred strains of mice
WR Atchley and WM Fitch
Extensive data on genetic divergence among 24 inbred strains of mice provide an opportunity to examine the concordance of gene trees and species trees, especially whether structured subsamples of loci give congruent estimates of phylogenetic relationships. Phylogenetic analyses of 144 separate loci reproduce almost exactly the known genealogical relationships among these 24 strains. Partitioning these loci into structured subsets representing loci coding for proteins, the immune system and endogenous viruses give incongruent phylogenetic results. The gene tree based on protein loci provides an accurate picture of the genealogical relationships among strains; however, gene trees based upon immune and viral data show significant deviations from known genealogical affinities.
======================
Science, Vol 255, Issue 5044, 589-592
Experimental phylogenetics: generation of a known phylogeny
DM Hillis, JJ Bull, ME White, MR Badgett, and IJ Molineux
Department of Zoology, University of Texas, Austin 78712.
Although methods of phylogenetic estimation are used routinely in comparative biology, direct tests of these methods are hampered by the lack of known phylogenies. Here a system based on serial propagation of bacteriophage T7 in the presence of a mutagen was used to create the first completely known phylogeny. Restriction-site maps of the terminal lineages were used to infer the evolutionary history of the experimental lines for comparison to the known history and actual ancestors. The five methods used to reconstruct branching pattern all predicted the correct topology but varied in their predictions of branch lengths; one method also predicts ancestral restriction maps and was found to be greater than 98 percent accurate.
==================================
Science, Vol 264, Issue 5159, 671-677
Application and accuracy of molecular phylogenies
DM Hillis, JP Huelsenbeck, and CW Cunningham
Department of Zoology, University of Texas, Austin 78712.
Molecular investigations of evolutionary history are being used to study subjects as diverse as the epidemiology of acquired immune deficiency syndrome and the origin of life. These studies depend on accurate estimates of phylogeny. The performance of methods of phylogenetic analysis can be assessed by numerical simulation studies and by the experimental evolution of organisms in controlled laboratory situations. Both kinds of assessment indicate that existing methods are effective at estimating phylogenies over a wide range of evolutionary conditions, especially if information about substitution bias is used to provide differential weightings for character transformations.
Application of the tested methodology:
Implications of natural selection in shaping 99.4% nonsynonymous DNA identity between humans and chimpanzees: Enlarging genus Homo
"Here we compare ≈90 kb of coding DNA nucleotide sequence from 97 human genes to their sequenced chimpanzee counterparts and to available sequenced gorilla, orangutan, and Old World monkey counterparts, and, on a more limited basis, to mouse. The nonsynonymous changes (functionally important), like synonymous changes (functionally much less important), show chimpanzees and humans to be most closely related, sharing 99.4% identity at nonsynonymous sites and 98.4% at synonymous sites. "
Mitochondrial Insertions into Primate Nuclear Genomes Suggest the Use of numts as a Tool for Phylogeny
"Moreover, numts identified in gorilla Supercontigs were used to test the human–chimp–gorilla trichotomy, yielding a high level of support for the sister relationship of human and chimpanzee."
A Molecular Phylogeny of Living Primates
"Once contentiously debated, the closest human relative of chimpanzee (Pan) within subfamily Homininae (Gorilla, Pan, Homo) is now generally undisputed. The branch forming the Homo and Pan lineage apart from Gorilla is relatively short (node 73, 27 steps MP, 0 indels) compared with that of the Pan genus (node 72, 91 steps MP, 2 indels) and suggests rapid speciation into the 3 genera occurred early in Homininae evolution. Based on 54 gene regions, Homo-Pan genetic distance range from 6.92 to 7.90×10−3 substitutions/site (P. paniscus and P. troglodytes, respectively), which is less than previous estimates based on large scale sequencing of specific regions such as chromosome 7[50]. "
Catarrhine phylogeny: noncoding DNA evidence for a diphyletic origin of the mangabeys and for a human-chimpanzee clade.
"The Superfamily Hominoidea for apes and humans is reduced to family Hominidae within Superfamily Cercopithecoidea, with all living hominids placed in subfamily Homininae; and (4) chimpanzees and humans are members of a single genus, Homo, with common and bonobo chimpanzees placed in subgenus H. (Pan) and humans placed in subgenus H. (Homo). It may be noted that humans and chimpanzees are more than 98.3% identical in their typical nuclear noncoding DNA and probably more than 99.5% identical in the active coding nucleotide sequences of their functional nuclear genes (Goodman et al., 1989, 1990). In mammals such high genetic correspondence is commonly found between sibling species below the generic level but not between species in different genera."
CONCLUSION:
This evidence lays out the results of employing a tested methodology on the question of Primate evolution. The same general criteria/methods have been used on nearly all facets of the evolution of living things.
As you like to pretend you understand all this, I eagerly await your analysis in which you point out the confirmation bias, circular reasoning, etc. that you pretend all actual science is based on.
But I will not be holding my breath. All hat, no cattle?
Upvote
0
If Cause-and-effect is so reliable to you, then what caused the First Cause?Oh no. Nonono. I already know my foundational presuppositions are not provable to you by your methods.
If Cause-and-effect isn't reliable to you, then First Cause certainly is not. No point in proceeding further.
Upvote
0
- Jan 9, 2018
- 3,132
- 871
- Country
- United States
- Gender
- Male
- Faith
- Christian
- Marital Status
- Single
It's called first cause for a reason. Lol.
Upvote
0
Hans Blaster
I march with Sherman
- Mar 11, 2017
- 23,201
- 17,247
- 55
- Country
- United States
- Gender
- Male
- Faith
- Atheist
- Marital Status
- Private
- Politics
- US-Democrat
I guess then that cause and effect isn't universal.
Upvote
0
- Jan 9, 2018
- 3,132
- 871
- Country
- United States
- Gender
- Male
- Faith
- Christian
- Marital Status
- Single
A first cause is necessarily outside of the universe.
Upvote
0
Hans Blaster
I march with Sherman
- Mar 11, 2017
- 23,201
- 17,247
- 55
- Country
- United States
- Gender
- Male
- Faith
- Atheist
- Marital Status
- Private
- Politics
- US-Democrat
A first cause is necessarily outside of the universe.
On what basis do you make this claim?
Upvote
0
No.A first cause is necessarily outside of the universe.
Nothing outside the observable universe (beyond the particle horizon) is causally connected with anything inside it.
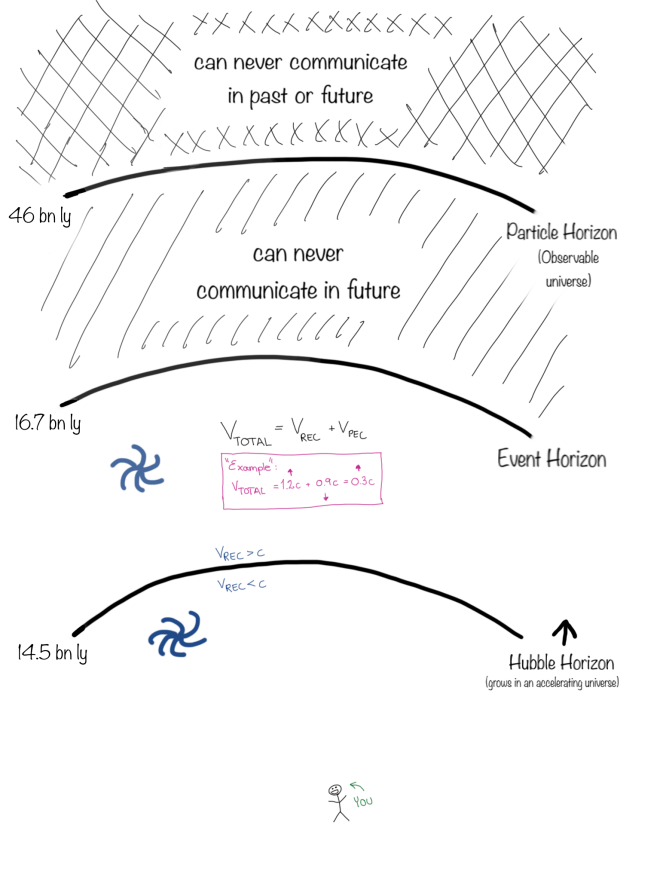
Upvote
0
- Jan 9, 2018
- 3,132
- 871
- Country
- United States
- Gender
- Male
- Faith
- Christian
- Marital Status
- Single
Everything we observe has a beginning a middle and an end.On what basis do you make this claim?
A first cause can't have a beginning or it is caused and not the first cause.
A first cause cannot be an effect.
But everything we can observe is.
Upvote
0
Hans Blaster
I march with Sherman
- Mar 11, 2017
- 23,201
- 17,247
- 55
- Country
- United States
- Gender
- Male
- Faith
- Atheist
- Marital Status
- Private
- Politics
- US-Democrat
Everything we observe has a beginning a middle and an end.
A first cause can't have a beginning or it is caused and not the first cause.
A first cause cannot be an effect.
But everything we can observe is.
So, basically, on no basis can you make that claim.
All you've got are pithy sounding nothings.
Upvote
0
- Jan 9, 2018
- 3,132
- 871
- Country
- United States
- Gender
- Male
- Faith
- Christian
- Marital Status
- Single
A first cause would not be subject to entropy. It is not caused therefore would be self existent, not subject to change or the law of physics. A first cause directly and indirectly caused everything that is.No.
Nothing outside the observable universe (beyond the particle horizon) is causally connected with anything inside it.

One can claim there is no such thing but that just removes that one from discussing what is intrinsic to a first cause.
Upvote
0
- Jan 9, 2018
- 3,132
- 871
- Country
- United States
- Gender
- Male
- Faith
- Christian
- Marital Status
- Single
So, basically, on no basis can you make that claim.
All you've got are pithy sounding nothings.
I must be doing well all you got is a pithy sounding nothing.All you've got are pithy sounding nothings
Edit. Minus the pith...
Last edited:
Upvote
0
Similar threads
- Replies
- 14
- Views
- 754
- Replies
- 1
- Views
- 150
- Replies
- 0
- Views
- 335
- Replies
- 50
- Views
- 2K

