I don't know about anyone else, but I don't see a problem here.
Retroelements and the human genome: New perspectives on an old relation
I wouldn't give it a second thought if not for the fact that these homology arguments are based on the appearance that the two genomes are virtually identical. I've run into this again and again with regards to the overall divergence. The two genomes diverge by 5% at least but the popular press continually tells us that we are 98% the same in our DNA, which is true, unless you count the indels.
The conclusion is the old saw that we share 98.5% of our DNA sequence with chimpanzee is probably in error. For this sample, a better estimate would be that 95% of the base pairs are exactly shared between chimpanzee and human DNA. In this sample of 779 kb, the divergence due to base substitution is 1.4%, and there is an additional 3.4% difference due to the presence of indels (
Divergence between samples of chimpanzee and human DNA sequences is 5%, counting indels)
I still wouldn't have a problem if not for the incessant claims that I'm in error. I'll get a flurry of posters who tell me I've been proven wrong or that I can't get my facts straight and yet none of you want to admit to the straightforward facts as they are clearly explained in the scientific literature.
I run into this every single time I try to discuss genomics on here. LM makes an argument based on an old Talk Origins argument saying ERVS are 1% of the Human Genome.
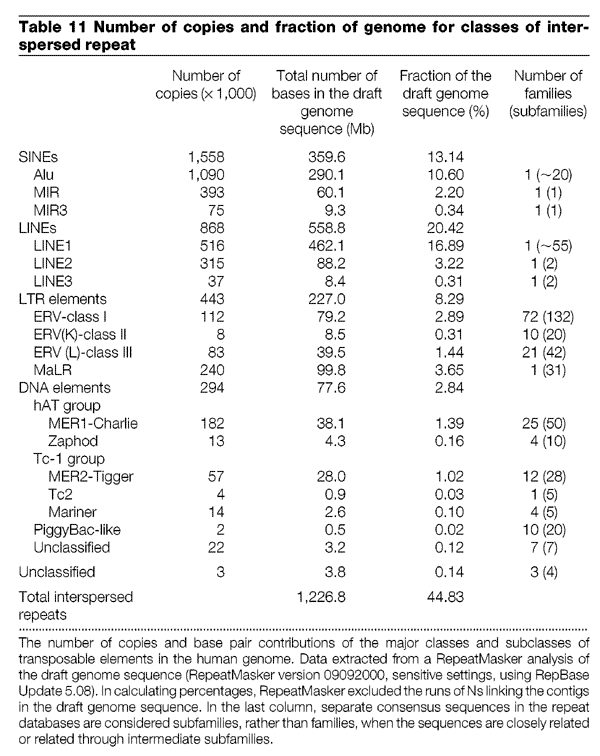
He eventually comes up with 4% based on the table in the Human Genome paper but the latest evidence I can find is that it's 8% of the Human genome overall. What is more those ERVs account for 7% of the divergence due to indels. He want's to argue that the ERVs in the respective genomes are 'virtually identical' which is odd given this:
According to this table, the chimp genome has 234 ERV class 1 elements not found in the human genome, and they total more than 1 million base pairs of sequence.
That's over a million base pairs of sequence that become a permanent part of the Chimpanzee genome as the result of germline invasions after the split. You really don't see a problem? Even thought the PtERV2 sequences are not as big,' the reconstructed subfamilies are too great (~8%) to have arisen by mutation since divergence from human.'.
There are obvious problems here that probably don't effect the overall homology argument much at all. It's simple enough, just make some adjustments in the overall percentages.
If all 234 were full length, and they run about 7 kb per repeat, that would be 1.6 million base pairs. That's 2% of the total contribution of ERV class 1 elements to the human or chimp genome. (234 is a much smaller fraction of the total number of ERVs in the genome, but that's not surprising: the recent, lineage-specific insertions are much more likely to still be full length, and therefore much longer than the typical ERV fossil, which just consists of the terminal repeat portion.)
Thank you! So what you are saying is that the divergent Class 1 ERVs make up 1.6 Mbps, a total of 2% in the respective genomes. I promise you I'm not trying to be pedantic here, I just want to clarify that this is a reliable statistic as an overall percentage.
So what was the problem again? And what does this have to do with the evidence that ERV insertion points were inherited?
Probably none really, I can make my standard arguments that the divergence is too great and the evolutionists can simple defend the view that their not. I really don't have a problem with that, it's what I would expect. If your convinced by the evidence of common ancestry that's fine. I would just like the facts used as evidence to be established without vacillating by millions of base pairs.
You've always been helpful with this sort of thing Steve. The debates often get heated which is part of the whole controversy between Creationism and Evolution anyway. I would appreciate just a little clarification on what the divergence actually is before I start fielding these incessant personal attacks.
That table also appears to be the source for all later statements of the fraction of ERVs in the human genome, which can be variously reported as 4.6%, 5-8% or "up to 8%". This is all really straight-forward.
That's not really all that straightforward. I fully appreciate that you might be looking at different things using different methods. Things are constantly being updated and revised so for the Human Genome paper to say ERVs make up 4.6% of the Human Genome and a report years later says that it's 8% it just means the estimate has been adjusted. If we are talking about less then 2% divergence overall it's not that big of a deal to explain it.
That's not what is going on here. Before the facts are established I'm getting these petty personal remarks about how I'm making glaring mistakes, lying about the facts and it's all just a figment of my 'perverse imagination'. Then when LM makes glaring errors he gets a pass. Hardly sounds objective to me.
Grace and peace,
Mark