Wiggle out?
Yep. That is exaclty what you do in the rest of your post.
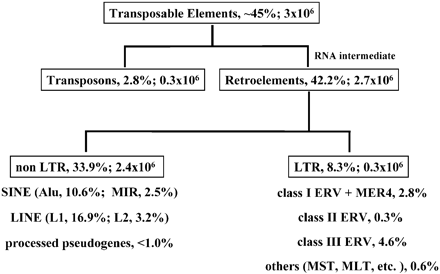
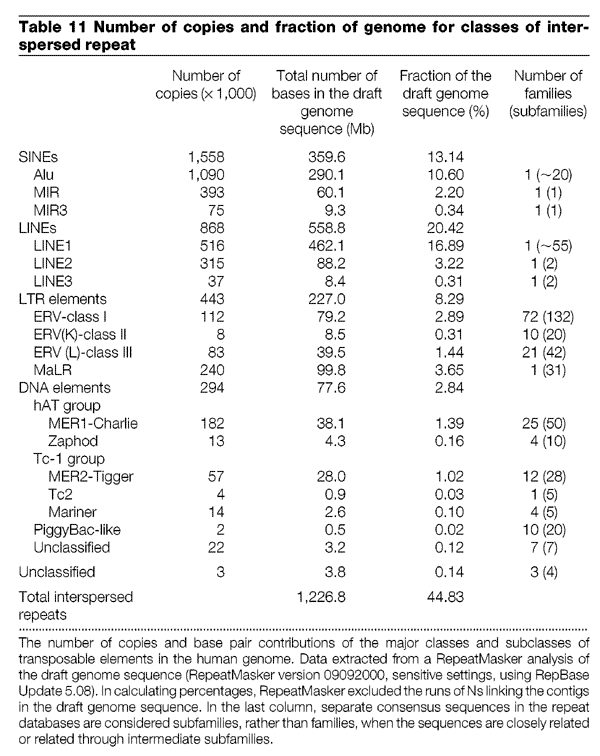
What the table says is that the ERV class 1 sequences are 79 million base pairs total, comprising under 3% of the entire sequence with 72 families. Ok, this is what we know about the Chimpanzee Genome:
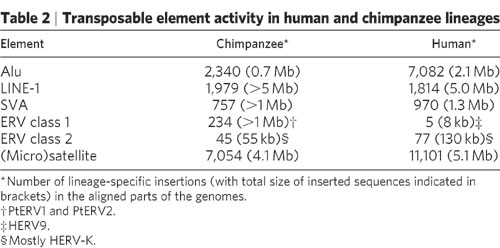

According to this table the Chimpanzee Genome has 234 ERV class 1 elements greater the 1 million base pairs (MPS). Do you see a problem here yet?
That is not what that chart says. That chart lists the ERV's that are specific to that lineage. Those are the ERV's that are not shared with humans. The other 200,000 are shared with humans which is why they are not listed in the chart for lineage specific transposable element activity. The ERV's listed for humans in that chart are the ERV's that have inserted into the human genome since the two lineages split. The ERV's listed for chimps are the ERV's that have inserted into the chimp genome since the two lineages split. That is why the chart lists separate lineages.
Want to guess how they determined which ERV's are lineage specific?
Just so we are on the same page, do you agree that there are ~200,000 ERV's in the human genome and that it is substantiated by Table 11 from the human genome paper.
Last edited:
Upvote
0