The second topic I would like to discuss from the Discovery Institute article by Jonathan M (JM hereafter) is Pterv insertions. As I will show, this argument actually blows up in the face of creationists. Remember what I said in a previous post about creationists trying to confuse the reader where it concerns species distribution and orthology? This is one of the best examples that one can find. Even more, it ends up supporting evolution in a way that completely contradicts the creationist argument.
The peer reviewed paper under discussion is this one:
Lineage-Specific Expansions of Retroviral Insertions within the Genomes of African Great Apes but Not Humans and Orangutans
The Discovery Institute webpage can be found here:
Do Shared ERVs Support Common Ancestry? - Evolution News & Views
It discusses the discovery of Pterv insertions which are found in the chimp and gorilla genomes, but curiously not in the human or orangutan genome. Pterv insertions are also found in other primate genomes. The theory of evolution would predict that with this species distribution, these insertions had to occur after the chimp and human lineages split. If they had occured in the common ancestor of chimps and gorillas, then humans should also have these insertions because humans share that same common ancestor. Humans don't have those insertions.
Since evolution predicts that these were independent insertions, it would also predict that given the number of insertions (about 300), we shouldn't find any or very, very few orthologous insertions.
This is where JM uses selective quotes to give the impression that there are 12 orthologous insertions shared by chimps and gorillas. It is in the grey quote box towards the bottom of the page that starts, "We performed two analyses to determine whether these 12 shared map intervals might indeed be orthologous. . ."
Strangely, or predictably, JM does not discuss the analyses that they used to determine if those 12 candidates are truly orthologous. A bit of background . . .
So why are there 12 candidates to begin with? From the real scientific paper:
WARNING, HEAVY SCIENCE CONTENT TO FOLLOW, SKIP TO LOWER SECTION IF YOU WISH .. .
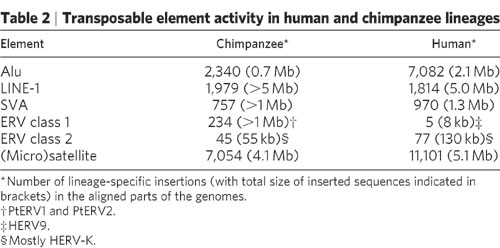
A total of 275 of the insertion sites mapped unambiguously to non-orthologous locations (Table 2), indicating that the vast majority of elements were lineage-specific (i.e., they emerged after the divergence of gorilla/chimpanzee and macaque/baboon from their common ancestor).
Within the limits of this BAC-based end-sequencing mapping approach, 24 sites mapped to similar regions of the human reference genome (approximately 160 kb) and could not be definitively resolved as orthologous or non-orthologous (Table S3). We classified these as ambiguous overlap loci (Figure 3). If all 24 locations corresponded to insertions that were orthologous for each pair, this would correspond to a maximum of 12 orthologous loci. [emphasis mine]
So what do they mean by a "BAC-based end-sequencing mapping approach". You first need to understand what a BAC clone is. A BAC clone is an E. coli clone that carries a plasmid with a big chunk of genomic DNA from the species being studied. They break the chimp, gorilla, or other genome down into chunks anywhere from 10,000 to 500,000 base pairs, and put them in a plasmid so that E. coli can store and replicate that DNA. When you sequence DNA, you need to start from known DNA. Since you know the DNA sequence of the plasmid, you can start your sequencing there and sequence into the big chunk of DNA. Most sequencing reads are about 1,000 base pairs, so you can get the sequence for the 1,000 base pairs at either end of the big chunk of genomic DNA. This allows you to know where that chunk belongs in the larger genome.
However, you only know the 1,000 base pairs at each end. This means that the ERV insertions detected in these BAC clones could be anywhere within that big chunk of DNA. All they know is that it is in there somewhere, but they don't know which base it is at. This is what they mean by "Within the limits of this BAC-based end-sequencing mapping approach". They only have the ends, and not the middle where the insertion is.
END OF HEAVY SCIENCE CONTENT, IF YOU SKIPPED THE MATERIAL, START HERE . . .
So as we can see, the BAC based method can not show us if these 12 candidates are truly orthologous. They can't show that the insertions are at the same base, only that they are within 10,000 or 100,000 bases of each other. What JM does not want you to know is that they used other methods to determine if these insertions were truly orthologous.
Here is the rather large quote that you can skip and read my summary below:
For the three intervals putatively shared between macaque and chimpanzee, we attempted to refine the precise position of the insertions by taking advantage of the available whole-genome shotgun sequences for these two genomes. For each of the three loci, we mapped the precise insertion site in the chimpanzee and then examined the corresponding site in macaque (
National Center for Biotechnology Information). In one case, we were unable to refine the map interval owing to the presence of repetitive rich sequences within the interval. In two cases, we were able to refine the map location to single basepair resolution (Figures S4 and S5). Based on this analysis, we determined that the sites were not orthologous between chimpanzee and macaque. It is interesting to note that this level of refined mapping in chimpanzee revealed 4- to 5-bp AT-rich target site duplications in both cases. These findings are consistent with an exogenous retrovirus source since proviral integrations typically target AT-rich DNA ranging from 4 to 6 bp in length [24]. Although the status of the remaining overlapping sites is unknown, these data resolve four additional sites as independent insertion events and suggest that the remainder may similarly be non-orthologous.
: IOW, for the insertions that they could confirm down to the base pair, they weren't orthologous, and they suspect the same will be true of the others. To claim that there are 12 orthologous Pterv insertions shared by chimps and gorillas based on BAC clones is a massive leap of faith.
But here is a more interesting aspect of this data, IMHO. ID/creationists will argue that it makes sense that ERV's are found in the same spot in the chimp and human genome because a common creator would use common building blocks. Since chimps and gorillas supposedly share a common creator, shouldn't we see the Pterv insertions shared in the same way that other ERV's are shared between humans and apes? I don't see why not.
What we have here is a set of DIFFERENT predictions made by the common creator group and the evolution group. The comon creator group makes the argument that Pterv insertions should be orthologous, just like the hundreds of thousands of ERV's that are orthologous between the chimp and human genome.
Evolutionists make the exact opposite argument based on the phylogeny of humans and apes. Evolutionists predict that they should NOT be orthologous because they would have to have been independent insertions that occurred after the human and chimp lineages split. This is based on a phylogeny, something that the common creator argument does not have.
So how do those predictions fare? The common creator argument is completely refuted, and the evolutionary prediction turns out to be completely right. Go figure.